Let us be honest, one of the reasons we use R Markdown to compile documents into PDF is the aesthetic pleasure provided by LaTeX. However, all the efforts can be ruined by wrong fonts in plots that are not the same as in the rest of the document. For example
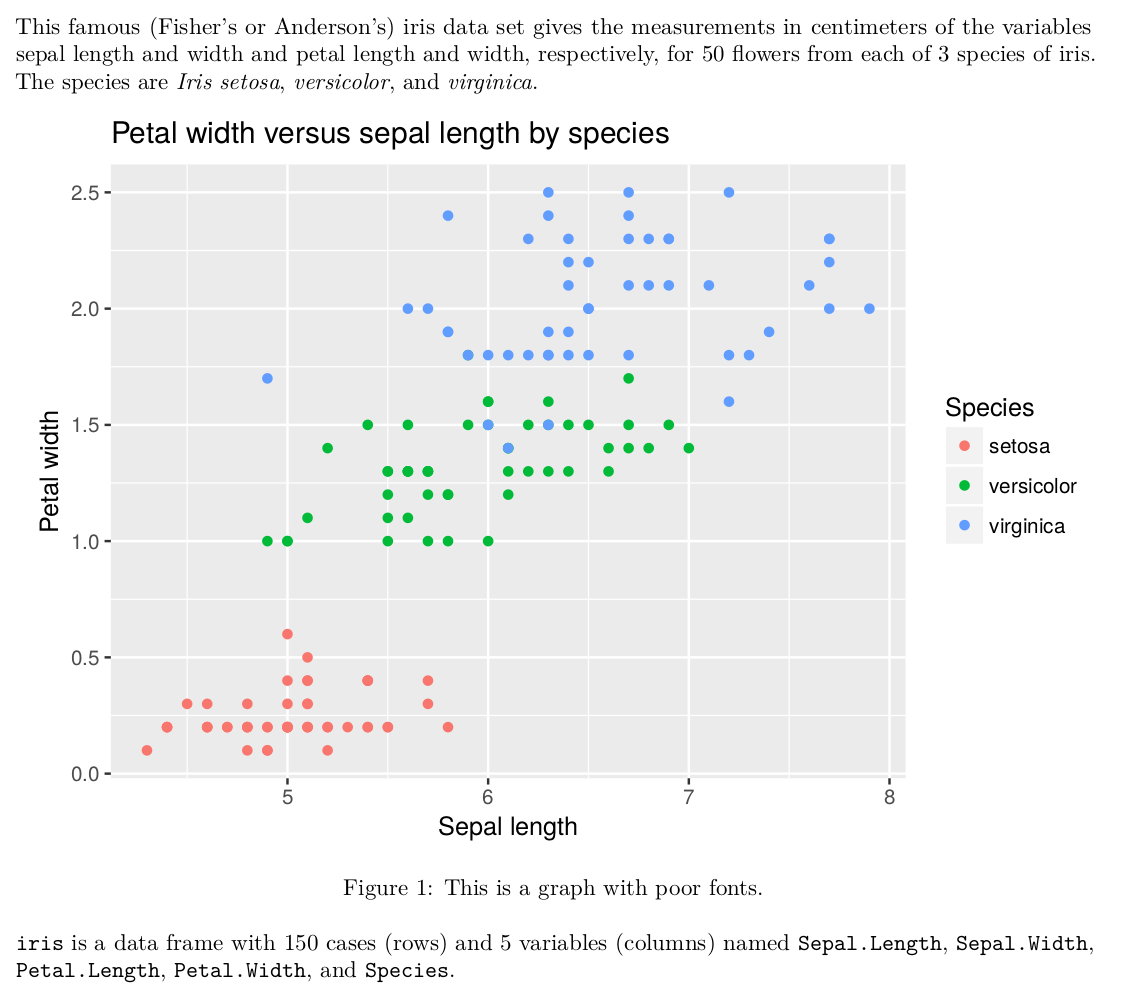
Well, these are some ugly fonts (not by itself, but in combination with the rest of the document). Would not it be much better to have something like this?
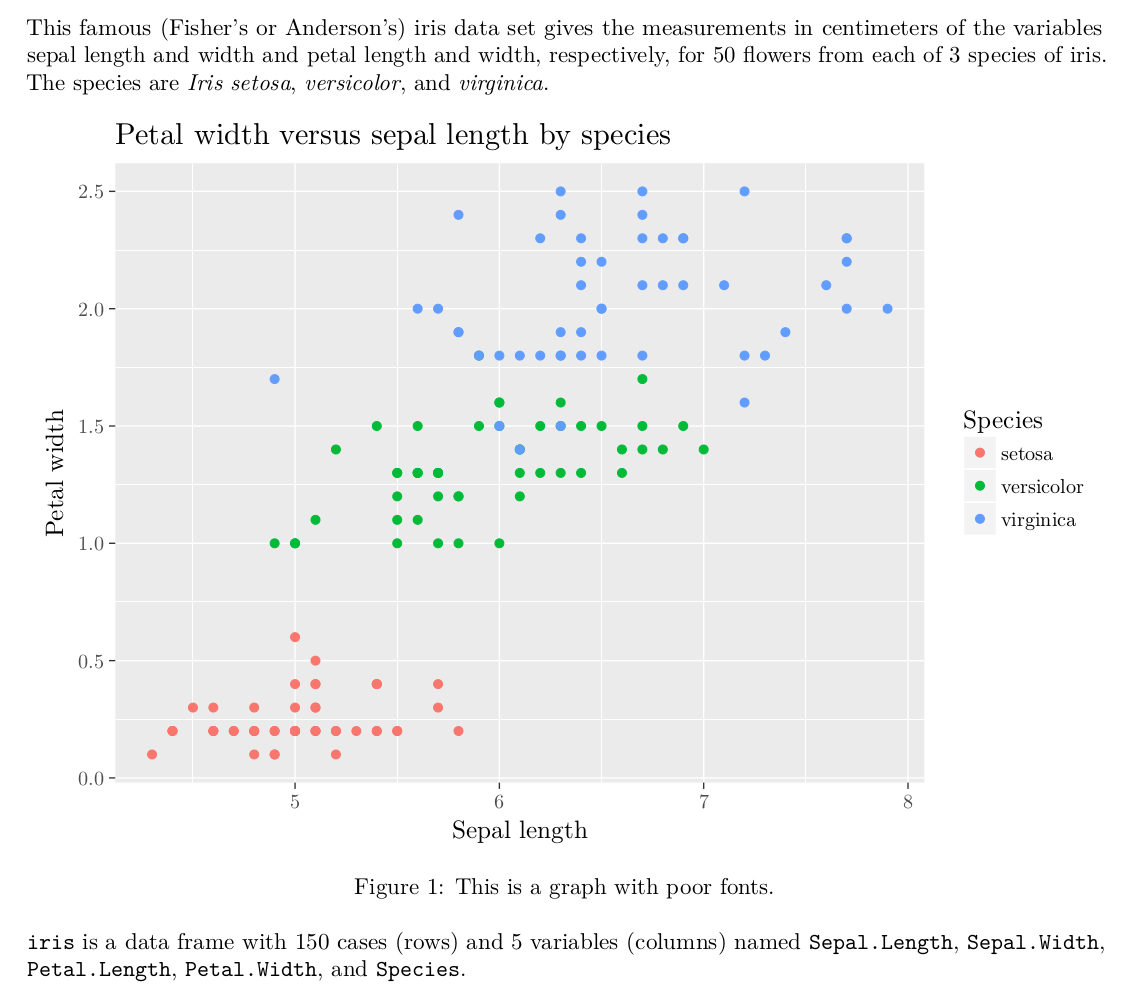
So how can we make the good ones? Well, the answer is easy: just let LaTeX render your images and it will take care of the rest.
Why LaTeX? In short, because it is a LaTeX engine that compiles a PDF. Remember
that for producing a PDF, knitr converts R Markdown to Pandoc Markdown, which
converts it to LaTeX, which and compiles it into PDF using a specified engine.
If you have not changed it, then it’s probably pdflatex, but that is not
important in absolute majority of cases.
Whenever we render a picture, for example a plot, using either the base plot
function or ggplot, we send it to a printing device. When it appears somewhere
in RStudio (highly likely that it appears in the bottom right corner), it means
that it was printed to the RStudioGD device. When we save it to a file, we
print it to, for example, a png or pdf device. And if we want to save it in
a format that is understood by LaTeX, we print with the device that produces the
desired output. Pictures in (La)TeX are compiled from
TikZ code. Thus, we need to print our
pictures to a tikz device. Luckily, someone (or rather Kirill Müller) was kind
enough to write a package that adds tikz device via
tikzDevice
package, which is availble on CRAN.
One might wonder: “But we do not specify a printing device when use R Markdown!”
This is partially true. If we do not specify a device in an R script, it will
attempt to print to the current active device, which is highly likely to be
RStudioGD. If we do not specify it in R Markdown, it will attempt to print to
its default device. There is an option for chunks, however, that allows
specifying the dev option, which stands for “device”. We can either set it
equal to "tikz" for all chunks by adding this option to the opts_chunk
knitr::opts_chunk$set(..., dev = "tikz")
or specify it for each chunk manually, like this
```{r, dev = "tikz"}
# Some plotting code goes here, e.g.
plot(rnorm(10), rnorm(10))
```Reproducible example
Reproducible example can be found here.
TL; DR
Make sure you have
tikzDevice
package, then either specify dev = "tikz" option for all chunks like this
knitr::opts_chunk$set(..., dev = "tikz")
or specify it for each chunk separately.